Genome analysis of enterobacteriaceae
Here is the summary of our last article, full article is available here : https://www.sciencedirect.com/science/article/pii/S0378113519312180
##BACKGROUND
Extended-spectrum-β-lactamases (ESBL) and plasmid-mediated cephalosporinases (pAmpC)-pro- ducing Enterobacteriaceae isolates are now reported worldwide in humans, animals, and in the environment. We identified the determinants of resistance to β-lactams and associated resistance genes as well as phylogenetic diversity of 53 ESBL- or pAmpC-producing Enterobacteriaceae isolated from dogs and cats in Europe.
##MATERIALS/METHODS
Of a collection of 842 Enterobacteriaceae isolates that were recovered in 2013 and 2014 from 842 diseased and untreated dogs and cats, for 242 ampicillin or amoxicillin resistant isolates (MIC ≥ 16 mg/L), cefotaxime (CTX) and ceftazidime (CAZ) MICs were determined. Isolates with CTX and/or CAZ MIC ≥ 1 mg/L (n = 63) were selected, and their genomes were fully sequenced using Illumina Technology. Genomic data were explored to identify the resistance determinants, the plasmid incompatibility groups, and the sequence types (STs). Plasmid location of bla ESBL and bla AmpC was evaluated for all isolates based on the co-localization of resistance and plasmid incompatibility group genes on the same contig. Phylogenetic trees were constructed using core-genome MLST
##RESULTS
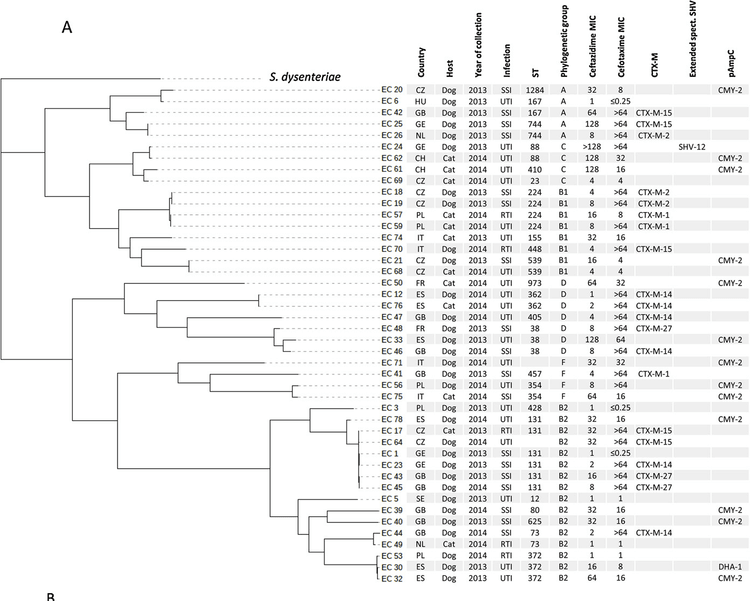
Of the 63 sequenced isolates, 53 isolates harbored a bla ESBL or bla AmpC gene. Ten CTX and/or CAZ non- wild type isolates had neither bla ESBL nor bla AmpC . Among the 63 isolates, 44 (69.8 %) were Escherichia coli, 11 (17.5 %) were Klebsiella pneumoniae, and 8 (12.7 %) were Proteus mirabilis. Fifty-one (80.9 %) isolates originated from dogs and 12 (19.1 %) from cats. Isolates were sampled from urinary tract (n = 36), skin and soft tissue (n = 22) and respiratory tract infections (n = 5). Thirty-two isolates (32/53, 60.4 %) carried bla ESBL genes, in- cluding bla CTX-M-15 (n = 12), bla CTX-M-14 (n = 6), bla CTX-M-1 (n = 5), bla CTX-M-2 (n = 3), bla CTX-M-27 (n = 3), bla SHV-28 (n = 4), bla SHV-12 (n = 2), and bla VEB-6 (n = 1). Four isolates of K. pneumoniae had both bla CTX-M-15 and bla SHV-28 . Twenty-one isolates (21/53, 39.6 %) carried genes encoding pAmpC, including bla CMY-2 (n = 19) and bla DHA-1 (n = 2). Thirteen E. coli isolates harbored both bla ESBL or bla AmpC genes and plasmids of incompatibility groups IncIB (9/13), IncI1 (8/13), and IncFII (6/13). In addition to the reduced susceptibility to CTX and/or CAZ, reduced susceptibility or evidence of acquired resistance to at least one other relevant class of antibiotics was observed for all 63 isolates. E. coli: isolates clustered in 23 STs, including B2 virulent clones from humans such as ST131 (n = 5), K. pneumoniae isolates mostly clustered in 3 STs: ST11 (n = 4), ST307 (n = 3), and ST16 (n = 2). Phylogenetic analysis identified the spread of E. coli ST131 bla CTX-M-27 , and of K. pneumoniae ST307 harboring bla CTX-M-15 and bla SHV-28 or ST11 bla CTX-M-15 .
##CONCLUSION
We report here a 6.3 % prevalence of ESBL/pAmpC producing Enterobacteriaceae in diseased dogs and cats. This EU survey confirms that dogs and cats can be infected with epidemic multidrug resistant clones that may also spread in humans.